AtaGenix Laboratories
AtaGenix Laboratories
Release time: 2025-06-20 View volume: 528
Project Snapshot — A rice biology team at the China National Rice Research Institute studying grain chalkiness needed recombinant CHALK10 (MBP-tag) and SD1 (GST-tag) proteins plus rabbit polyclonal antibodies for co-IP and ubiquitination assays. AtaGenix delivered E. coli-expressed proteins and validated antibodies that confirmed the CHALK10–SD1 interaction and its role in GA-mediated starch metabolism — published in Plant Communications (2025).
Grain chalkiness in rice (Oryza sativa) negatively impacts grain quality, affecting texture, milling resilience, and consumer preference. A 2025 study published in Plant Communications (DOI: 10.1016/j.xplc.2025.101354) by researchers at the State Key Laboratory of Rice Biology and Breeding identified CHALK10, an F-box protein that negatively regulates grain chalkiness by mediating ubiquitination and degradation of SEMIDWARF-1 (SD1) via the 26S proteasome pathway. CHALK10 interacts with SD1 to modulate gibberellin (GA) levels and starch metabolism, thereby affecting endosperm development and chalkiness formation.
The research team required: (1) high-purity recombinant CHALK10 and SD1 proteins with affinity tags for co-immunoprecipitation (co-IP) assays to validate their physical interaction in rice endosperm; (2) rabbit polyclonal antibodies against both CHALK10 and SD1 for immunoblotting in ubiquitination and degradation studies; (3) proteins and antibodies compatible with in vitro ubiquitination assay workflows; and (4) validated reagent quality sufficient for a high-impact publication in plant molecular biology.
This plant biology project presented specific challenges:
AtaGenix designed an integrated protein + antibody workflow for this plant biology application:
The high-purity recombinant proteins and specific polyclonal antibodies enabled the research team to confirm the CHALK10–SD1 physical interaction via co-IP and demonstrate that CHALK10 mediates SD1 ubiquitination and 26S proteasomal degradation. This reduces bioactive GA levels, alters starch biosynthesis gene expression during grain filling, and ultimately decreases grain chalkiness. The study established CHALK10 as a novel molecular target for improving rice grain quality through breeding, with direct implications for global food security and consumer acceptance of rice varieties.

Figure 1. Graphical overview of the CHALK10–SD1 axis. The F-box protein CHALK10 targets SD1 for ubiquitination and 26S proteasomal degradation, modulating gibberellin levels and starch metabolism to regulate rice grain chalkiness.
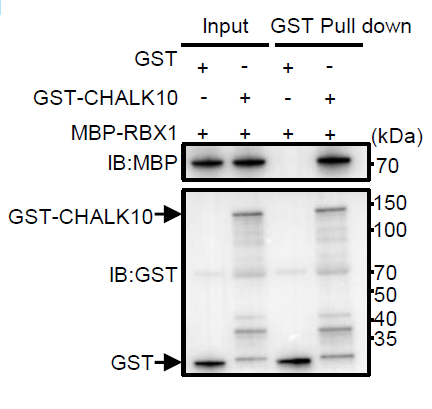
Figure 2. Co-immunoprecipitation and immunoblot validation of the CHALK10–SD1 interaction using AtaGenix-produced recombinant proteins and rabbit polyclonal antibodies. Specific detection confirmed direct protein-protein interaction in rice protoplast extracts.
About AtaGenix
AtaGenix provides one-stop customized support from protein/antibody design to functional validation, covering recombinant protein expression (E. coli, yeast, insect, mammalian), custom polyclonal and monoclonal antibodies, and multi-platform validation (WB, co-IP, ELISA, ubiquitination assays) — tailored for plant molecular biology and beyond. Learn more at www.atagenix.com.
This case study is based on a published research collaboration. Results may vary depending on target protein, expression system, and experimental conditions. All proprietary client information is subject to NDA. Reference: Plant Communications. 2025. DOI: 10.1016/j.xplc.2025.101354
Need co-IP-grade recombinant proteins and matched antibodies for plant signaling, ubiquitination, or protein-protein interaction studies? AtaGenix delivers from construct design to assay-validated reagents.
Explore E. coli Expression ServicePolyclonal Antibody | Peptide Synthesis | Yeast Expression | Mammalian Expression
Contact Us
+86-27-6552-3339
info@atagenix.com
Building C, R & D Building, No. 666, Shendun 4th Road, Donghu New Technology Development Zone, Wuhan

