AtaGenix Laboratories
AtaGenix Laboratories
Release time: 2026-04-01 View volume: 227
March has been marked by frequent research breakthroughs. AtaGenix continues to empower collaborative teams with its core technologies, helping cutting-edge life science research reach new heights. This month, multiple collaborative achievements have been published in prestigious international journals such as Cancer Research, Nature Communications, and Nature Sensors. The studies span multiple disease areas including colorectal cancer, lung adenocarcinoma, and endometrial cancer, showcasing both the breadth and depth of life science exploration. With its one-stop professional technical services providing full support, AtaGenix lays a solid foundation for the efficient translation of research outcomes and continues to inject strong momentum into global scientific innovation.
Title: METTL3 Methylation Induces Decay of Endogenous Retroelement Transcripts to Promote Tumor Immune Evasion
Journal: Cancer Research
Impact Factor: 16.6
Affiliation: Huazhong University of Science and Technology

Colorectal cancer (CRC) is prone to recurrence and its immunosuppressive microenvironment hinders anti-tumor immunity, limiting the effectiveness of current treatments. The study found that dimethylation of METTL3 at the K513 site is associated with CRC progression and recurrence. SETD1A catalyzes methylation at this site, enhancing the binding of METTL3 to SAM and increasing RNA m⁶A modification. This in turn suppresses the expression of endogenous retroelements (EREs), blocks type I interferon responses, and promotes tumor immune evasion. Fluorouracil can induce E2F4 autoregulation, activating SETD1A-mediated METTL3 methylation. Targeting the E2F4/SETD1A/METTL3 axis or blocking METTL3 K513 methylation can restore anti-tumor immunity, and when combined with immune checkpoint inhibitors, it significantly inhibits tumor growth. This study reveals a novel mechanism of CRC immune evasion and provides new targets and strategies for CRC treatment.
AtaGenix customized a rabbit polyclonal antibody against METTL3 K513me2 for this research. It specifically detects the dimethylation status of the K513 site on the METTL3 protein, providing a direct detection tool to validate the association between METTL3 K513 methylation and CRC progression/recurrence, and supporting key conclusions such as SETD1A-catalyzed methylation at this site and METTL3 methylation regulating downstream signaling pathways.
Title: The transcription factor EHF promotes the maturation and immunosuppression of conventional dendritic cells
Journal: Nature Communications
Impact Factor: 15.7
Affiliation: Guangdong Provincial People's Hospital

The transcriptional regulatory mechanisms underlying the immunosuppressive function of dendritic cells (DCs) remain unclear, while CCR7⁺ conventional dendritic cells (cDCs) play a critical role in immune tolerance. This study identified ETS homologous factor (EHF) as a key transcription factor regulating cDC maturation and immunosuppression. Mice with EHF deficiency showed enhanced resistance to autoimmune diseases, infections, and tumors. EHF-deficient cDCs promoted Th1 and Th17 CD4⁺ helper T cell responses both in vitro and in vivo. EHF can be activated by TLR7/8/9 and inhibited by TLR3, GM-CSF, and IFNγ. It directly regulates the transcription of genes such as Ccr7, Cd200, Cd274, Irf4, and Rel, promoting the expression of CCR7, CD200, and PD-L1 while suppressing IRF4 expression. Additionally, EHF is highly enriched in CCR7-high DCs in both humans and mice. This study reveals a conserved transcriptional program for cDC immunosuppression and provides new targets for the treatment of autoimmune diseases, cancer, and infectious diseases.
AtaGenix provided an anti-EHF monoclonal antibody for the study. It specifically recognizes the EHF protein and served as a specific probe for CUT&TAG analysis, directly supporting the validation of EHF’s transcriptional regulation of target genes and clarifying the direct binding relationship between EHF and genes such as Ccr7 and Cd200. The antibody was also used in Western blotting and other experiments to verify the expression patterns of EHF across different tissues and cells, ensuring the reliability of the results regarding EHF expression and function.
Title: A cell cycle-dependent transition of acetylation to phosphorylation regulates timely centrosome maturation
Journal: Nature Communications
Impact Factor: 10.4
Affiliation: School of Medicine, University of Electronic Science and Technology of China

Centrosomes must mature during the G2/M phase to ensure bipolar spindle assembly and chromosome segregation. PLK1 is a key kinase in this process, but how the cell cycle regulates its timely recruitment remains unclear. This study found that the ubiquitin E3 ligase RNF40 localizes to the centrosome and is phosphorylated by CDK1 at T529/T557 sites during G2/M phase. This phosphorylation enables RNF40 to bind PLK1 and promote its recruitment, forming a CDK1-RNF40-PLK1 cascade regulatory mechanism. During interphase, RNF40 is acetylated by PCAF at K517/K562 sites. A transition from acetylation to phosphorylation occurs from late G2 to M phase, which is a prerequisite for cascade activation. Persistent acetylation or phosphorylation defects disrupt PLK1 localization, microtubule nucleation, and bipolar spindle assembly, leading to mitotic catastrophe. Furthermore, RNF40 dysregulation is associated with colorectal cancer, and targeting this cascade may enhance chemotherapy efficacy. This study reveals a novel cell cycle-dependent mechanism for centrosome maturation and provides new targets for cancer therapy.
AtaGenix prepared four specific antibodies for this study: rabbit polyclonal antibodies against pT529-RNF40, pT557-RNF40, acK517-RNF40, and acK562-RNF40. These antibodies precisely detect the phosphorylation or acetylation status of specific sites on RNF40, providing direct evidence for validating RNF40 modification characteristics. They ensured the reliability of core data regarding RNF40 modification sites, modification timing, and functional relevance, forming an important foundation for elucidating the CDK1-RNF40-PLK1 cascade and the biological significance of the RNF40 modification switch.
Title: HE4 drives PD-L1 expression in myeloid cells via IFN-γR-JAK-STAT3 signaling to promote tumor immune evasion
Journal: Cell Reports Medicine
Impact Factor: 10.6
Affiliation: School of Medicine, University of Electronic Science and Technology of China
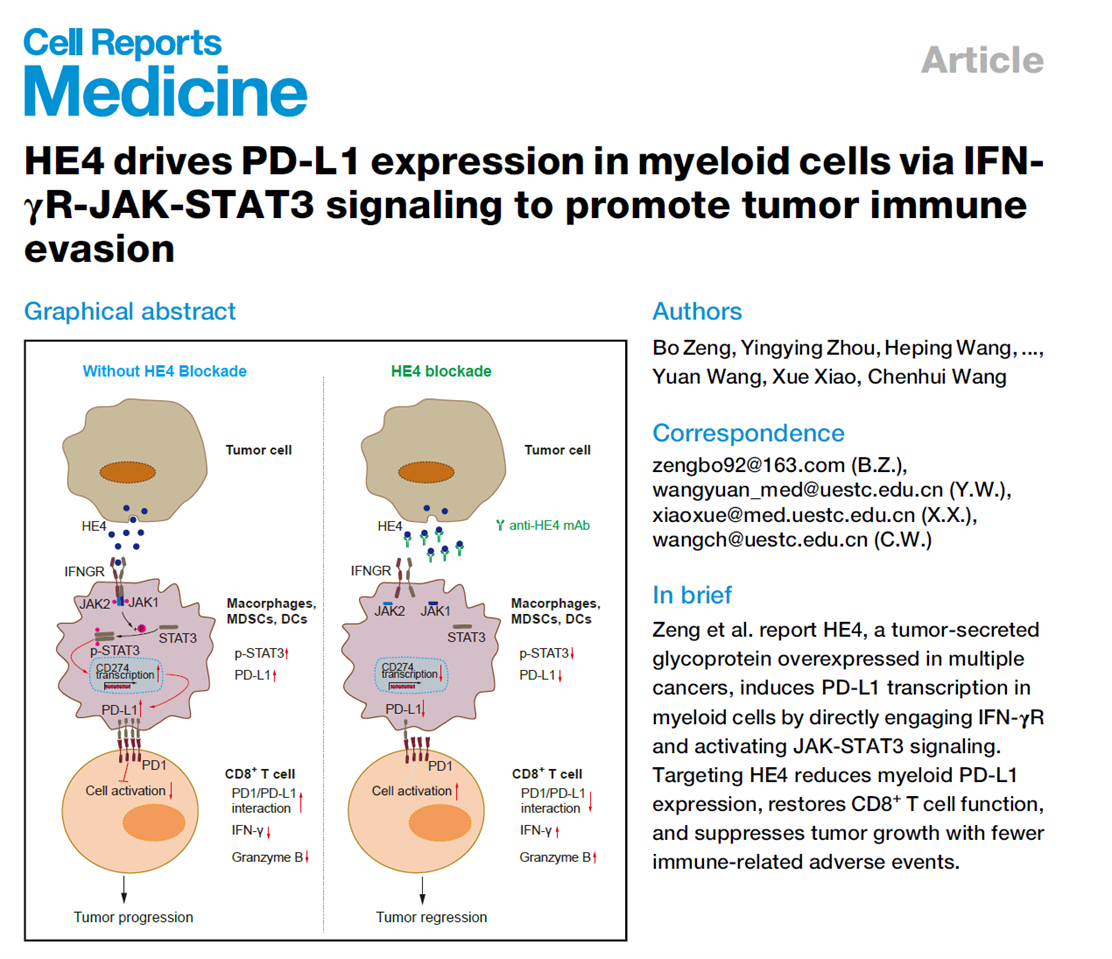
Clinical application of PD-1/PD-L1 immune checkpoint inhibitors is limited by low response rates and immune-related adverse events. The regulatory mechanisms of PD-L1 expression on myeloid cells in the tumor microenvironment remain unclear. This study found that tumor-secreted HE4 directly binds to IFN-γR on myeloid cells, activating the JAK-STAT3 signaling pathway, transcriptionally upregulating PD-L1 expression, suppressing CD8⁺ T cell function, and promoting tumor immune evasion. HE4 competitively binds to IFN-γR with IFN-γ, and high concentrations of HE4 in the tumor microenvironment can antagonize IFN-γ-mediated STAT1 activation. The developed anti-HE4 monoclonal antibody significantly reduced PD-L1 expression on myeloid cells, restored CD8⁺ T cell activity, inhibited tumor growth in multiple tumor models, and caused far fewer immune-related adverse events than PD-1 inhibitors. Clinical data showed that high HE4 expression can predict response to PD-1 inhibitors in lung adenocarcinoma patients, making it a potential therapeutic target and predictive biomarker.
AtaGenix prepared rat anti-mouse HE4 monoclonal antibody and mouse anti-human HE4 monoclonal antibody for this study. These antibodies were used to screen clones capable of specifically blocking HE4-induced PD-L1 upregulation, validating the critical role of HE4 in PD-L1 regulation. They were also employed in in vivo and in vitro anti-tumor experiments, confirming that HE4 neutralization can reverse the immunosuppressive tumor microenvironment. These antibodies serve as core reagents for evaluating the safety and efficacy of HE4 blockade therapy.
Title: Oriented nanobody–field-effect transistor interfaces enable ultrasensitive cancer biomarker detection
Journal: Nature Sensors
Impact Factor: —
Affiliation: China University of Geosciences

Early screening for endometrial cancer lacks efficient and portable methods. Traditional detection approaches suffer from high cost and long turnaround times. Although field-effect transistor (FET) biosensors hold great potential, their clinical translation is hindered by probe instability, non-specific adsorption, and Debye shielding. To address this, the research team developed the SNAP-FET platform, which combines genetic code expansion and click chemistry to achieve site-specific oriented immobilization of nanobodies, paired with an indium oxide transducer. They also integrated a portable device called ENDOCARE. The platform demonstrated ultra-high sensitivity, specificity, and stability in cell, animal model, and clinical samples, with results highly consistent with ELISA. The ROC curve AUC values for detecting 18 endometrial cancer patients and 11 healthy individuals reached 0.8434 (HE4) and 0.8131 (CA125), respectively. This study provides a new strategy for the clinical translation of FET biosensors and lays a technical foundation for point-of-care early diagnosis of tumors.
AtaGenix undertook the construction and screening of the CA125 nanobody phage display library for this study. The natural VHH phage display library (1.51 × 10¹⁰ colony-forming units) constructed by AtaGenix served as the core basis for screening CA125-specific nanobodies. After three rounds of panning and monoclonal screening, a high-affinity CA125-targeting nanobody NT2 was successfully obtained. It became the key recognition element for CA125 detection in the SNAP-FET platform, enabling multiplex detection of two core markers for endometrial cancer. This provided an important probe for validating the universality of the site-specific modification method and was a key component in demonstrating the broad applicability of the SNAP-FET platform.
Title: A proliferative subpopulation of coelomocytes in sea cucumber is identified and characterized by single-cell RNA-seq after evisceration
Journal: Fish and Shellfish Immunology
Impact Factor: 3.9
Affiliation: Yancheng Institute of Technology

Regeneration research is a global frontier in science. As a representative echinoderm with extremely strong regenerative capacity, the sea cucumber can rapidly regenerate coelomocytes after evisceration, but the underlying recovery mechanisms remain unclear. Using single-cell RNA sequencing, this study analyzed coelomocytes from the coelom and water vascular system of the sea cucumber Apostichopus japonicus and identified 19 subpopulations. The results showed that after evisceration, the proportion of subpopulation 10 in both the coelom and water vascular system increased by 15-fold and 10-fold, respectively. This subpopulation exhibits proliferative characteristics. Coelomic subpopulation 10 can directly differentiate into subpopulations 0, 4, and 5, while water vascular subpopulation 10 requires dedifferentiation before differentiating into subpopulation 9 (immune-related cells). The genes Rplp0 and piwi play key regulatory roles in this process. The study provides a comprehensive resource for the molecular classification of sea cucumber coelomocytes and reveals the dynamic changes in composition and function of coelomocytes following evisceration.
AtaGenix provided custom antibody preparation service for NLRC3 in the experiment. Since NLRC3 serves as a marker gene for subpopulation 9 and lacked suitable sequences for in-house antibody production, AtaGenix synthesized peptides from its highly conserved regions and prepared a specific primary antibody. In immunocytochemical staining, this antibody successfully clarified the morphological characteristics of subpopulation 9 cells (diameter 5–6 μm, spherical shape, moderate nucleus-to-cytoplasm ratio, with positive signals appearing as large granules in the cytoplasm). It provided key experimental evidence for validating the immune functional properties of subpopulation 9 and clarifying its differentiation relationship with subpopulation 10.
Thank you for your attention and support!
AtaGenix will continue to provide high-quality one-stop professional technical services including custom antibodies and protein expression, supporting more scientific breakthroughs.
For collaboration inquiries, please contact:
Email: info@atagenix.com
Phone: +86-027-65523339
Contact Us
+86-27-6552-3339
info@atagenix.com
Building C, R & D Building, No. 666, Shendun 4th Road, Donghu New Technology Development Zone, Wuhan

