AtaGenix Laboratories
AtaGenix Laboratories
Release time: 2025-12-16 View volume: 228
Streptococcus pyogenes secretes two key IgG-targeting enzymes: the protease IdeS (cleaves the hinge region into F(ab')₂ and Fc) and the glycosidase EndoS (hydrolyzes the Fc N-glycan). A Nature Communications study resolved the crystal structures of both enzyme–IgG1 Fc complexes, revealing the molecular basis of substrate-specific recognition. Clinical applications now span Guillain-Barré syndrome, AAV gene therapy antibody clearance, and anti-GBM renal disease.
IdeS, first reported in 1999, is a cysteine endopeptidase with highly specific IgG cleavage and no activity against IgM, IgA, IgD, or IgE. It is FDA-approved for preconditioning in hypersensitized kidney transplant patients. EndoS is widely used in antibody glycosylation engineering. The Nature Communications study (DOI: 10.1038/s41467-022-35340-z) established structural models for enzyme-specific IgG recognition, providing theoretical support for next-generation enzyme development.
The study addressed IgG Fc self-crystallization by identifying a key E382–R255 salt bridge in the crystal lattice. Three "Fx" mutants (E382A/S/R) disrupted this contact, with E382S significantly reducing crystallization efficiency and altering the space group — clearing obstacles for enzyme–Fc complex structural analysis and providing a general strategy for studying IgG-binding proteins.
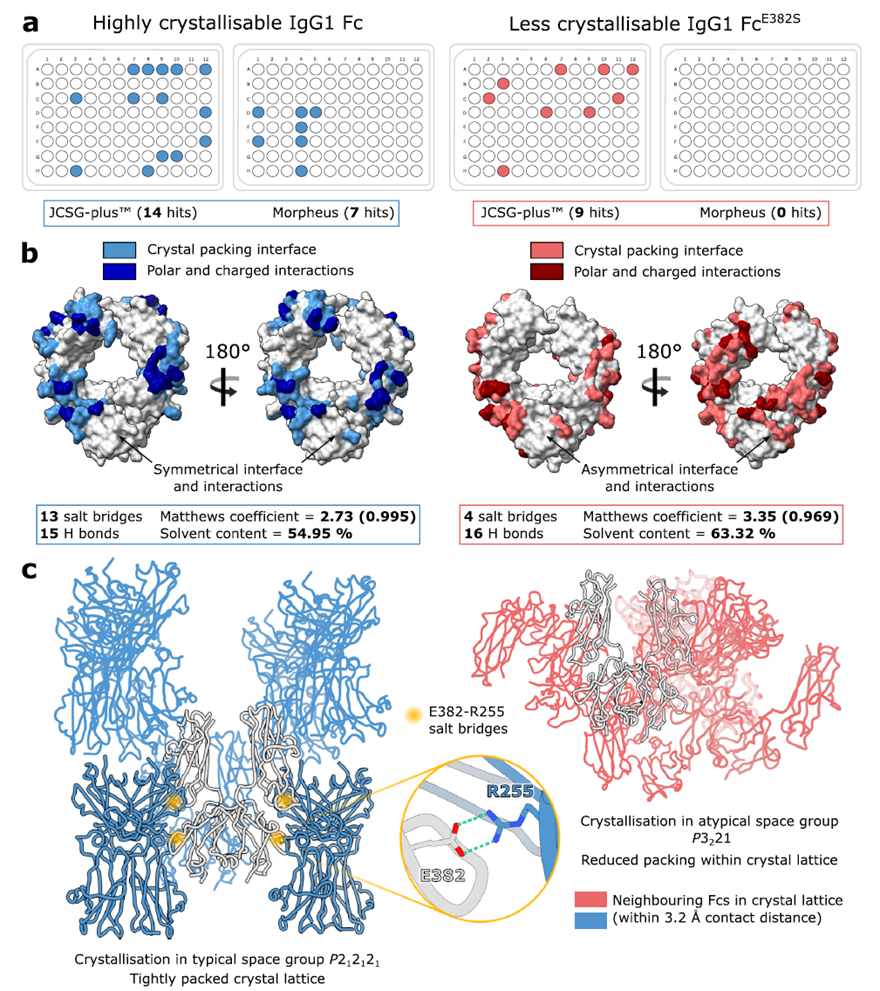
Fig. 1. Crystal packing in wild-type and "less crystallizable" IgG1 Fc fragments. Adapted from Sudol et al., Nat Commun 2022.
Catalytically inactive mutants (IdeSC94A, EndoSD233A/E235L) were mixed with Fx variants at 1:1 molar ratio, purified by SEC, and crystallized. X-ray crystallography resolved the 3D structures of both IdeS–IgG1 FcE382A and EndoS–IgG1 FcE382R complexes, clarifying interaction interfaces and key catalytic residues.
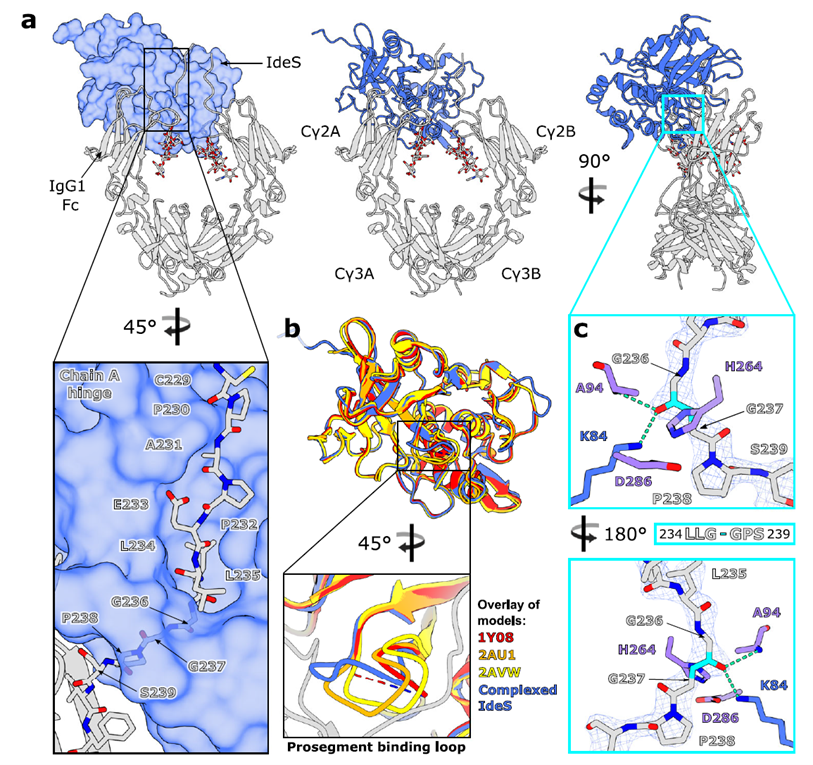
Fig. 2. Crystal structure of IgG1 FcE382A–IdeSC94A complex. Adapted from Sudol et al., Nat Commun 2022.
IdeS binds asymmetrically to the Cγ2 domain and lower hinge of IgG1 Fc in a 1:1 monomeric stoichiometry, "clamping" the hinge to form the active site chamber. The β7–β8 loop curls upward to mediate substrate entry. Dual interfaces with Fc (A-chain 1,392 Ų; B-chain 804.7 Ų) achieve recognition via hydrogen bonds and hydrophobic interactions. The binding site overlaps with Fcγ receptors, inhibiting Fc-mediated effector functions.
EndoS binds IgG1 Fc via a "V"-shaped structure, causing the N-glycan to "flip" from between Cγ2 domains to the exterior and embed into the glycosidase domain. The carbohydrate-binding module (CBM) binds the Cγ2–Cγ3 interface via hydrophobic interactions of W803. This forms a 2:2 complex (two EndoS molecules binding the two Fc chains separately), supporting efficient glycan hydrolysis.
IdeS significantly reduced nerve fiber damage in rabbit GBS models, lowered mortality, and blocked nerve injury progression by degrading pathogenic antibodies (Experimental Neurology, 2017). A 2024 multicenter clinical trial of 30 adult GBS patients receiving single-dose IdeS + IVIg showed 37% recovering independent walking within one week, with recovery 6 weeks earlier than conventional treatment and a 6.4-fold higher likelihood of independent walking at week 1.
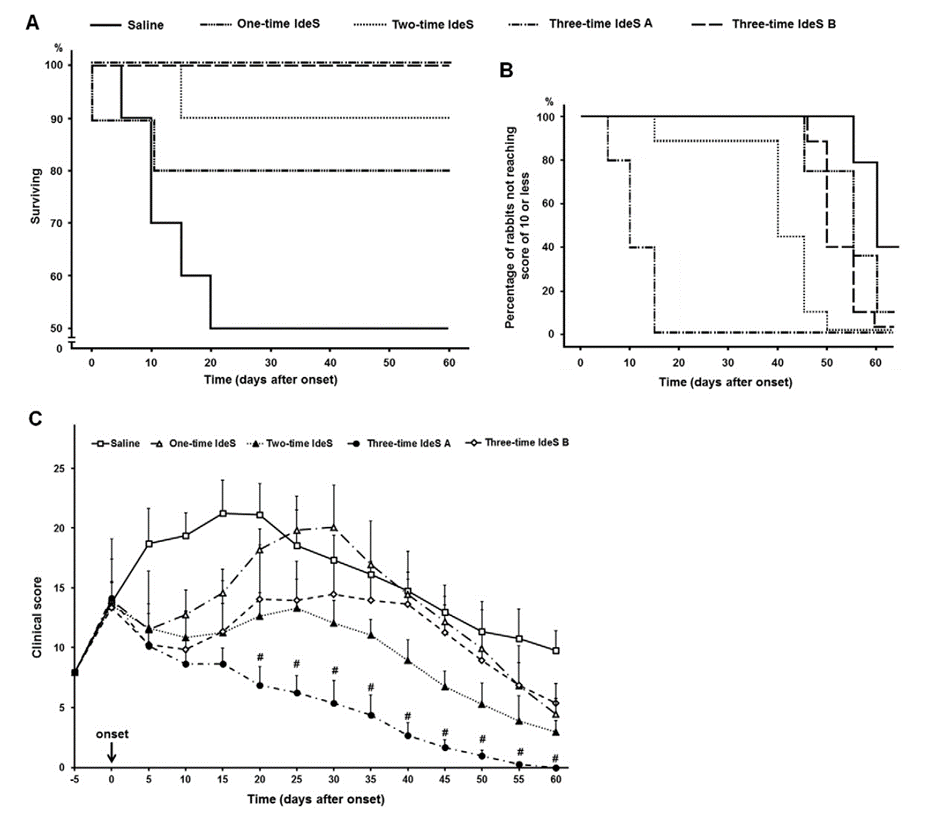
Fig. 3. IdeS significantly improves clinical symptoms in a rabbit AMAN model. Adapted from Wang et al., Exp Neurol 2017.
Pre-existing anti-AAV neutralizing antibodies block liver transduction and prevent vector readministration. A Nature Medicine study (2020) showed IdeS efficiently cleaves anti-AAV IgG in vitro and in vivo, enabling effective liver gene transfer in passively immunized mice and non-human primates. IdeS acts as an "antibody clearance tool" that directly degrades existing antibodies without broad immunosuppression.
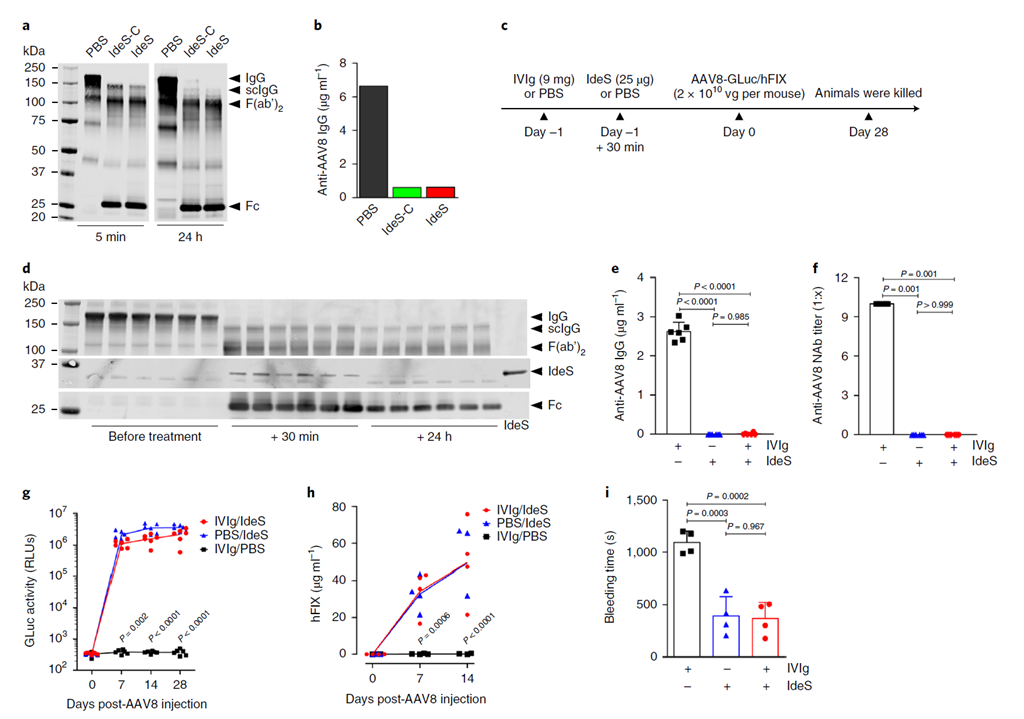
Fig. 4. IdeS degrades anti-AAV antibodies and enables liver transduction in IVIg-immunized mice. Adapted from Leborgne et al., Nat Med 2020.
The GOOD-IDES Phase II study (JASN) enrolled 15 severe anti-GBM patients (eGFR ≤15 mL/min/1.73 m²) receiving a single 0.25 mg/kg IdeS dose alongside standard immunosuppression. At 6 months, most achieved dialysis-independent renal function with persistent anti-GBM antibodies below toxic thresholds and significant relief of hematuria and proteinuria.
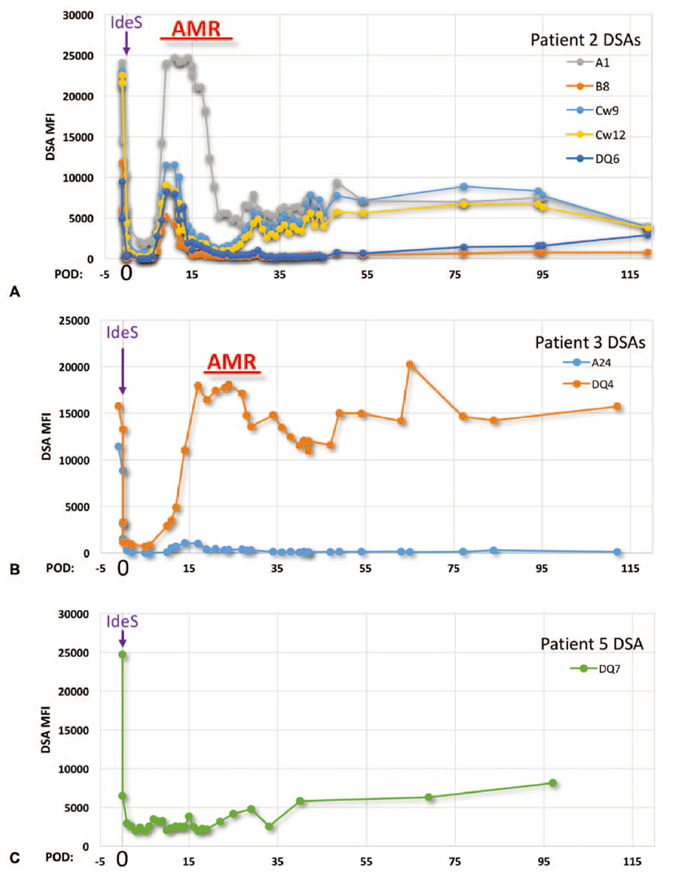
Fig. 5. Temporal trends of donor-specific antibody (DSA) levels in select patients. Adapted from JASN.
Clinical Treatment: Structure-optimized IdeS variants for expanded autoimmune indications (SLE, vasculitis) and antibody-mediated rejection prevention across organ transplantation.
Biopharmaceutical: Structural characterization of therapeutic antibodies, Fc fusion protein functional analysis, and ADC development — IdeS can inactivate competing serum IgG to enhance therapeutic antibody in vivo efficacy.
Gene Therapy: "Enzyme + vector" combination therapies resolving neutralizing antibody barriers for AAV, lentiviral, and other vector systems across hereditary diseases and oncology.
Anti-Infection: Design of IdeS-resistant biologics based on IdeS–IgG binding sites for treating severe streptococcal infections including necrotizing fasciitis.
AtaGenix Recombinant IdeS Protease
Catalog No. ATE00010 — Purity >95%, activity up to 40 U/μL, free of animal-derived contaminants. Suitable for therapeutic antibody structural characterization, ADC development, gene therapy preclinical experiments, and more.
| Catalog No. | Product Name |
|---|---|
| JN999012 | IdeS Protease (IgG specific) |
| JN865012 | Endo S2 / Endo-β-N-acetylglucosaminidase |
1. Wang Y, et al. Exp Neurol. 2017;291:134–140. DOI: 10.1016/j.expneurol.2017.02.010
2. Lonze BE, et al. Ann Surg. 2018;268(3):488–496. DOI: 10.1097/SLA.0000000000002924
3. Leborgne C, et al. Nat Med. 2020;26(7):1096–1101. DOI: 10.1038/s41591-020-0911-7
4. Sudol ASL, et al. Nat Commun. 2022;13:7801. DOI: 10.1038/s41467-022-35340-z
Need high-purity IdeS/EndoS proteases for antibody engineering, gene therapy, or structural biology? AtaGenix delivers recombinant enzyme tools with >95% purity and validated activity.
Talk to Technical SupportResponse within 24 hours
Contact Us
+86-27-6552-3339
info@atagenix.com
Building C, R & D Building, No. 666, Shendun 4th Road, Donghu New Technology Development Zone, Wuhan

