AtaGenix Laboratories
AtaGenix Laboratories
Release time: 2025-08-25 View volume: 876
Project Snapshot — A 2021 Small Methods study mapped immunogenic peptides across the SARS-CoV-2 spike protein, identified three key RBM epitopes with diagnostic potential (AUC ~0.98–1.0), and isolated patient-derived neutralizing monoclonal antibodies using a 8.7×109 ScFv phage library. AtaGenix provided antibody library construction, phage display screening, and antibody expression services that were instrumental in the discovery workflow.
Based on: Li et al., Small Methods, 2021 (DOI: 10.1002/smtd.202100058)
The SARS-CoV-2 spike (S) protein is the primary target of neutralizing antibodies and the basis of most COVID-19 vaccine strategies. Within its receptor-binding domain (RBD), the receptor-binding motif (RBM) directly contacts human ACE2 and is therefore a hotspot for protective immunity. But in 2021, the precise immunogenic epitopes within the spike — and which of those epitopes actually elicit neutralizing antibodies in patients — remained incompletely defined. Closing this gap was essential for rational vaccine design, diagnostic development, and therapeutic antibody discovery.
The research team designed a two-library strategy that combined epitope mapping and antibody discovery in a single integrated workflow:
Approximately 15 peptides across the spike were recognized by patient sera, but three RBM epitopes stood out: S431–454, S470–486, and S501–515. All three are surface-exposed and divergent from SARS-CoV and MERS-CoV, indicating SARS-CoV-2 specificity.
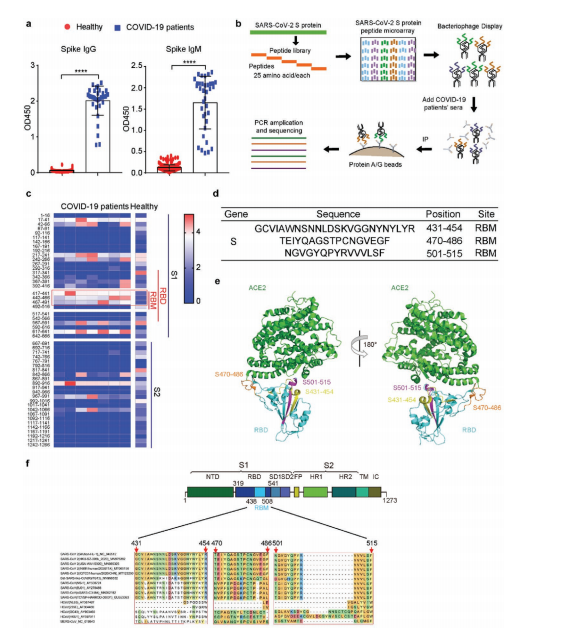
Figure 1. Phage display peptide library screening identified ~15 immunogenic spike peptides. Three epitopes within the RBM (S431–454, S470–486, S501–515) showed the strongest and most consistent antibody reactivity across patient sera.
ELISA testing across 43 confirmed patients, 117 suspected patients, and 38 healthy controls showed that antibodies against these three peptides — particularly IgG — distinguished COVID-19 patients from healthy individuals with near-perfect sensitivity and specificity.
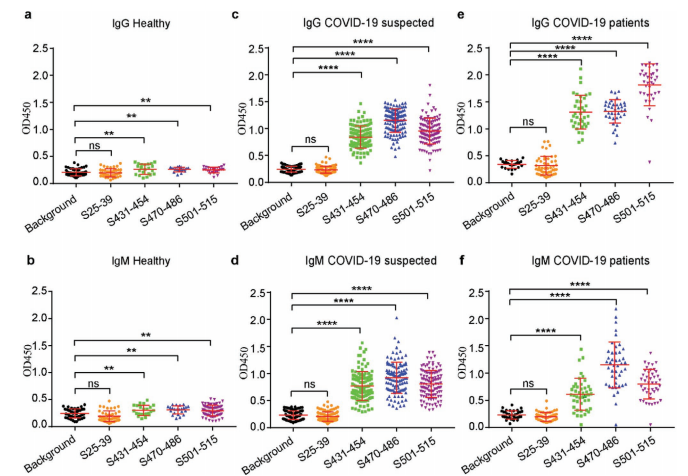
Figure 2. Antibody levels (IgM and IgG) against the three RBM peptides were significantly elevated in confirmed and suspected COVID-19 patients compared to healthy controls, demonstrating diagnostic utility with AUC values approaching 1.0.
Synthetic peptides were tested for their ability to block viral entry. S470–486 inhibited pseudovirus infection in vitro, and — crucially — mice immunized with S470–486 generated neutralizing antibodies effective against authentic SARS-CoV-2. The other two epitopes (S431–454, S501–515) elicited binding antibodies but not neutralizing ones, establishing S470–486 as the immunodominant protective epitope within the RBM.
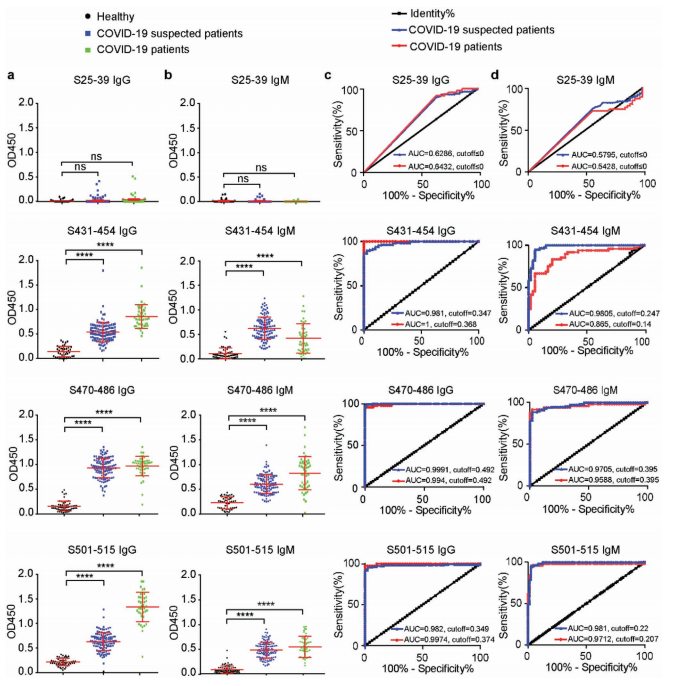
Figure 3. S470–486 blocked pseudovirus entry in vitro and elicited neutralizing antibodies in immunized mice, while S431–454 and S501–515 did not produce neutralizing responses — establishing S470–486 as the key protective epitope.
From the 8.7×109-clone ScFv phage library, two monoclonal antibodies targeting S470–486 were isolated. One clone (R3P1-F8) demonstrated potent neutralization against both pseudovirus and authentic SARS-CoV-2, proving that functional therapeutic antibodies can be rapidly obtained directly from patient immune repertoires using phage display — without the need for hybridoma or single B cell approaches.
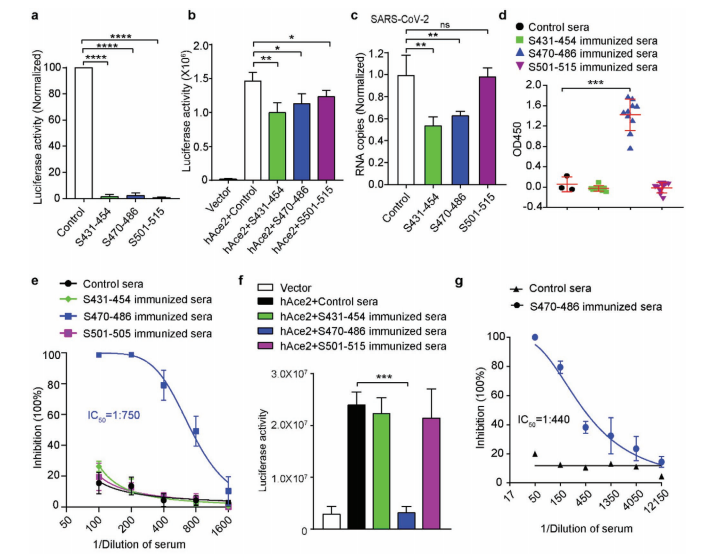
Figure 4. Human monoclonal antibodies isolated from a COVID-19 patient ScFv phage library bound the S470–486 epitope and neutralized SARS-CoV-2 in both pseudovirus and authentic virus assays. Clone R3P1-F8 showed the strongest neutralization activity.
AtaGenix provided critical technical services that underpinned the discovery workflow:
This study demonstrated a complete workflow from epitope discovery to therapeutic antibody isolation in a single campaign: map immunogenic peptides → validate diagnostic utility → identify the neutralizing epitope → isolate patient-derived mAbs targeting that epitope. The S470–486 region emerged as a high-value target for both diagnostics and therapeutics. Critically, the entire discovery cycle was accelerated by phage display technology — bypassing the slower hybridoma and single B cell approaches — with AtaGenix's library construction and screening capabilities enabling the research team to achieve 109-scale coverage that would have been difficult to build in an academic setting alone.
Li L, Gao M, Li J, et al. Methods to Identify Immunogenic Peptides in SARS-CoV-2 Spike and Protective Monoclonal Antibodies in COVID-19 Patients. Small Methods. 2021;5:2100058. DOI: 10.1002/smtd.202100058
Why This Matters
Phage display remains one of the fastest paths from patient sample to therapeutic antibody candidate — but only if the library is large and diverse enough. Building a 109-clone ScFv library from patient PBMCs requires specialized infrastructure that most academic and early-stage biotech labs don't maintain in-house. This collaboration demonstrates how partnering with AtaGenix for library construction and screening gave the research team access to pharma-grade phage display capabilities, compressing what would otherwise be a 6–12 month in-house effort into weeks — and producing a publication-quality dataset with real neutralizing antibodies at the end.
This case study is based on a published research collaboration. Results may vary depending on target, library diversity, and patient sample quality. All proprietary client information is subject to NDA.
Need phage display library construction, epitope mapping, or patient-derived antibody discovery? AtaGenix provides end-to-end phage display services from library building through functional antibody validation.
Discuss Your Antibody Discovery ProjectResponse within 24 hours
Contact Us
+86-27-6552-3339
info@atagenix.com
Building C, R & D Building, No. 666, Shendun 4th Road, Donghu New Technology Development Zone, Wuhan

