AtaGenix Laboratories
AtaGenix Laboratories
Release time: 2024-11-05 View volume: 946
Overview — Three case studies showcasing the depth of AtaGenix's E. coli expression capabilities: (1) systematic 32-condition optimization with post-purification His-tag removal, (2) periplasmic expression to rescue a disulfide-bonded antibody fragment from inclusion bodies, and (3) inclusion body refolding to recover functional protein from insoluble aggregates. Each case demonstrates a different problem-solving strategy within the same prokaryotic platform.
Client (NDA) | Structural Biology Program
Challenge: A target protein expressed poorly in standard BL21/IPTG/37°C conditions, with most product partitioning into inclusion bodies. The client also needed the final protein tag-free for crystallography — even a small His-tag can interfere with crystal packing.
Solution: AtaGenix ran a systematic optimization screen across multiple variables simultaneously:
| Variable | Conditions Tested |
|---|---|
| Host Strains | T7E, BL21(DE3), C41(DE3), ArcticExpress |
| Induction Temperature | 16°C (overnight) / 37°C (rapid) |
| Induction Time | 4 h (37°C) / 16 h (16°C) |
| Inducer | IPTG (multiple concentrations) |
| Culture Media | LB / Auto-induction medium |
The optimal condition (low-temperature induction in ArcticExpress with auto-induction medium) shifted the majority of expression to the soluble fraction. After Ni-NTA purification, the His-tag was cleanly removed using 3C protease, and the tag-free protein was polished by a subtractive Ni-NTA step (cleaved protein in flow-through, uncleaved + free tag retained on column).
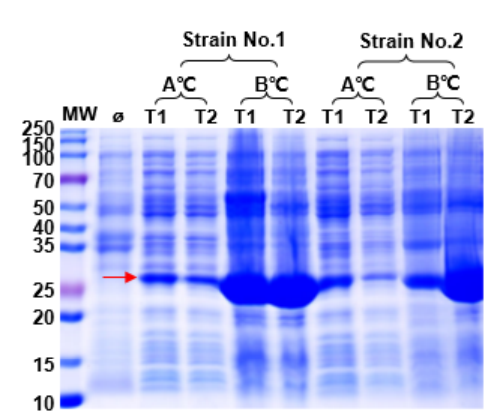
Figure 1. Expression optimization screen. Varying host strain, temperature, induction time, and media composition identified conditions that dramatically improved soluble protein yield and reduced inclusion body formation.
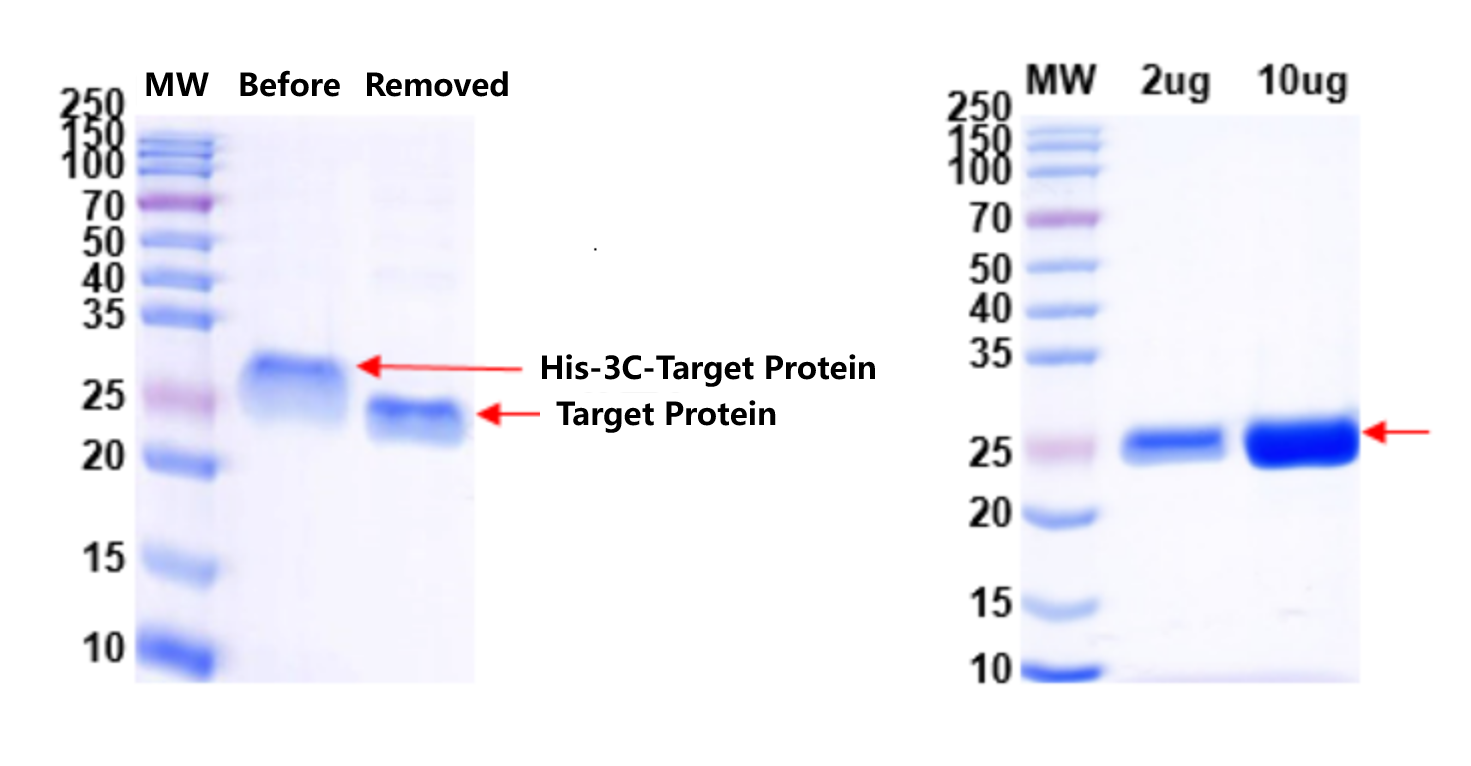
Figure 2 & 3. His-tag removal and final QC. Left: 3C protease efficiently cleaved the His-tag with near-complete conversion. Right: QC of tag-free protein confirmed target integrity, high purity, and absence of uncleaved precursor — ready for crystallization trials.
Client (NDA) | Antibody Engineering Program
Challenge: An antibody fragment (Fab or scFv) contains disulfide bonds essential for its structure. When expressed in the E. coli cytoplasm — a reducing environment — these bonds cannot form, causing the protein to misfold and aggregate into inclusion bodies. The client needed correctly folded, functional antibody fragment without the time and risk of refolding.
Solution: AtaGenix redirected expression to the E. coli periplasm — the oxidizing compartment between the inner and outer membranes. A signal peptide (e.g., pelB or OmpA) directed the nascent polypeptide across the inner membrane into the periplasmic space, where the DsbA/DsbB oxidase system catalyzes disulfide bond formation. The periplasmic fraction was selectively released by osmotic shock (no full cell lysis), reducing host cell protein contamination and simplifying purification.
Result: SDS-PAGE under both reducing and non-reducing conditions confirmed correctly folded antibody fragment with intact disulfide bonds. The non-reduced sample migrated as a compact band (oxidized), while the reduced sample showed the expected upward shift (linearized). No inclusion body processing or refolding was required.
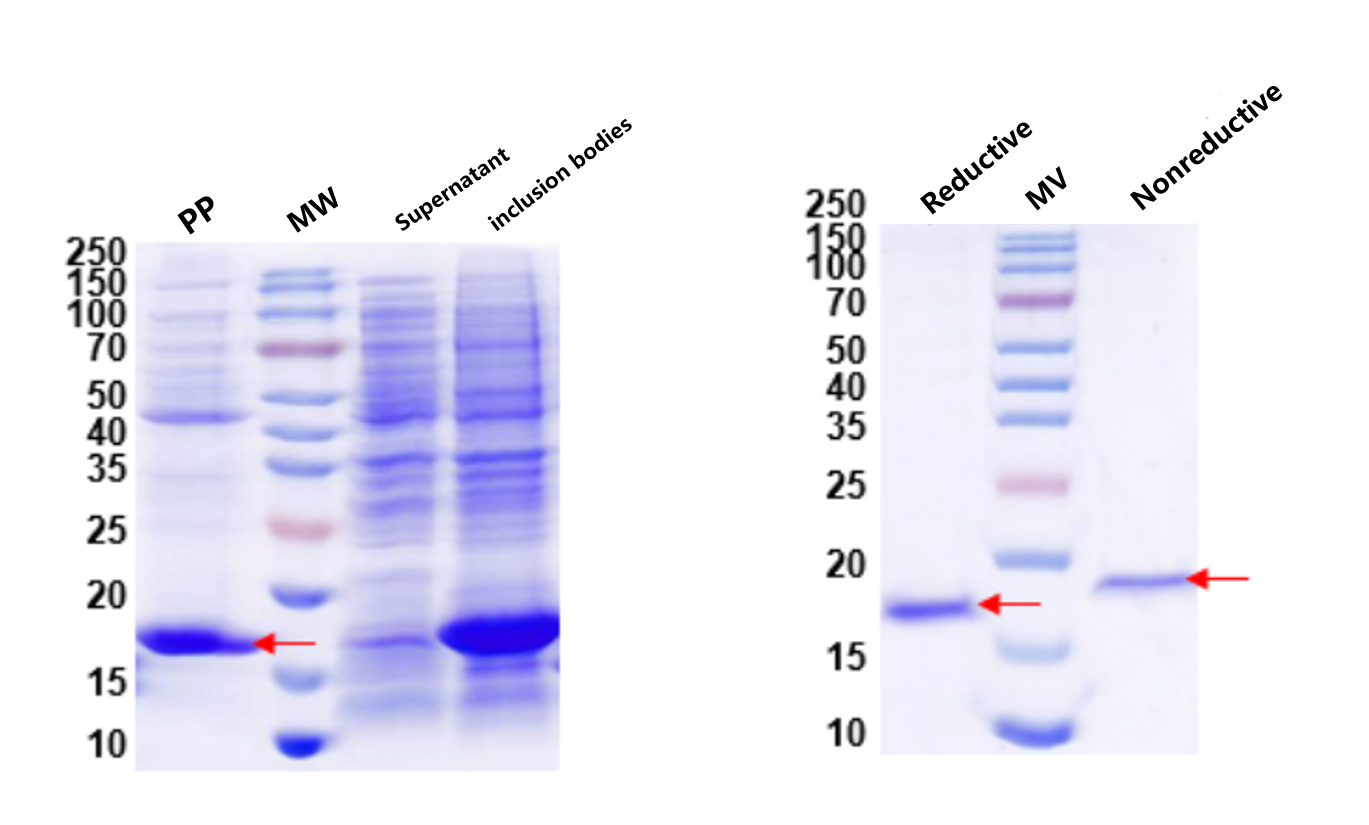
Figure 4 & 5. Periplasmic expression and QC. Left: Expression analysis showing soluble antibody fragment in the periplasmic fraction (Lane PP). Right: Reduced vs. non-reduced SDS-PAGE confirms intact disulfide bonds — the hallmark of correctly folded antibody fragments. MW: protein marker.
Client (NDA) | Enzyme Research Program
Challenge: A target protein expressed exclusively as inclusion bodies in E. coli, regardless of strain, temperature, or inducer optimization. Periplasmic expression was not an option (the protein was too large for efficient translocation). The client needed functional protein for activity assays — meaning the refolded product had to recover enzymatic activity, not just become soluble.
Solution: AtaGenix isolated inclusion bodies, solubilized them in 8 M urea with reducing agent, and applied a gradient dialysis refolding protocol — stepwise urea removal with controlled redox conditions (GSH/GSSG) to allow progressive refolding. The refolding buffer composition, protein concentration, dilution rate, and temperature were optimized for this specific target. The refolded soluble fraction was separated from residual aggregates by centrifugation and further purified by affinity chromatography.
Result: SDS-PAGE confirmed conversion from insoluble inclusion bodies to a clean soluble band at the expected molecular weight. The client subsequently confirmed that the refolded protein retained enzymatic activity in their functional assay — the ultimate test of successful refolding.
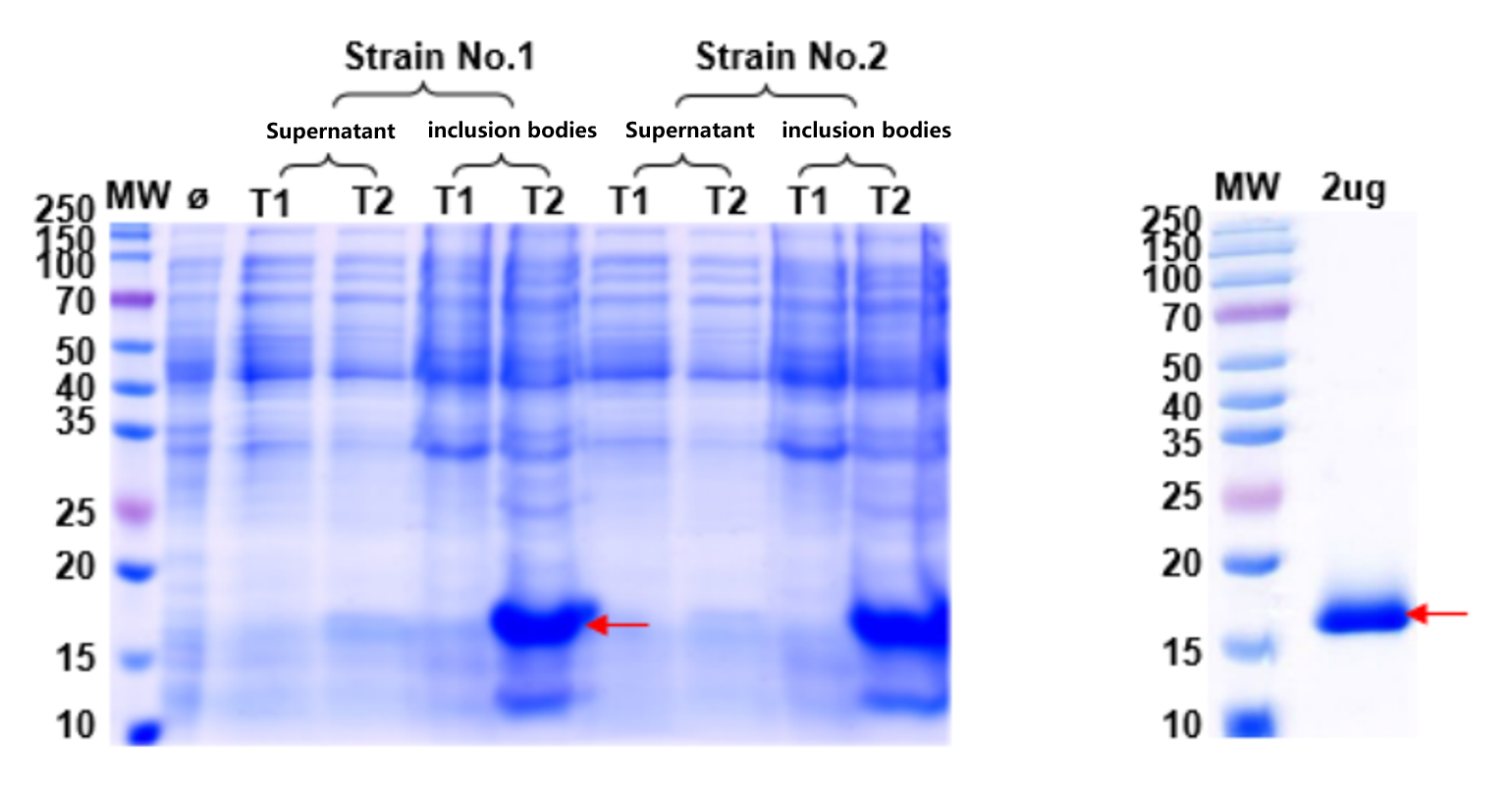
Figure 6 & 7. Left: Expression optimization confirmed the target protein forms inclusion bodies across all conditions tested. Right: After urea solubilization and gradient dialysis refolding, SDS-PAGE shows successful conversion to soluble, purified protein — subsequently confirmed as enzymatically active by the client.
| Problem | Strategy | Best For |
|---|---|---|
| Low solubility at 37°C | Multi-condition optimization | Proteins that can be soluble with the right strain/temp/media combination |
| Disulfide bonds needed | Periplasmic expression | Antibody fragments (Fab, scFv), small disulfide-rich proteins |
| Obligate inclusion bodies | Inclusion body refolding | Large or complex proteins that won't fold in any cytoplasmic condition |
| Tag interferes with function | Enzymatic tag removal (3C, TEV) | Structural biology, functional assays, in vivo studies |
Why This Matters
E. coli is the fastest and most cost-effective expression system — when it works. The real question is what to do when it doesn't. Most CROs offer a single approach: "we'll try BL21 at 16°C, and if it doesn't work, switch to mammalian." AtaGenix exhausts the full E. coli toolkit first — strain screening, temperature/media optimization, periplasmic redirection, inclusion body refolding, and tag removal — before recommending a more expensive eukaryotic system. This saves clients both time and money, because when E. coli can deliver, nothing is faster or cheaper.
These case studies represent anonymized project outcomes. Results may vary depending on target protein, construct design, and project scope. All client information is subject to NDA.
Struggling with insoluble protein, misfolded antibody fragments, or need tag-free protein for structural studies? AtaGenix will find the right E. coli strategy for your target — or recommend the best alternative if prokaryotic expression isn't suitable.
Discuss Your Expression ChallengeResponse within 24 hours
Contact Us
+86-27-6552-3339
info@atagenix.com
Building C, R & D Building, No. 666, Shendun 4th Road, Donghu New Technology Development Zone, Wuhan

