AtaGenix Laboratories
AtaGenix Laboratories
Release time: 2026-04-03 View volume: 287
Affinity tags such as His, GST, MBP, and SUMO greatly improve the yield and solubility of recombinant proteins. However, the tags themselves often interfere with the target protein’s structure, function, or downstream applications. Therefore, precise tag removal has become a critical step. Enzymatic cleavage is currently the most reliable method due to its high specificity. This article summarizes the advantages and limitations of commonly used endoproteases and exoproteases, and integrates strategies for solubility tag design and cleavage.
Affinity tags facilitate purification but can alter protein folding, oligomeric state, or immunogenicity. Enzymatic reagents offer sufficient specificity and are widely used for tag removal. Traditional serine proteases (such as enterokinase, thrombin, and Factor Xa) exhibit high activity but carry a significant risk of non-specific cleavage. In contrast, viral cysteine proteases (TEV and HRV 3C) have low k_cat values, resulting in extremely high specificity with almost no reported off-target cleavage. SUMO proteases operate differently, relying on recognition of the substrate’s overall folded structure to achieve “zero-residue” cleavage.
Detergent sensitivity studies show that thrombin is completely tolerant to 94 different detergents, making it the preferred choice for membrane protein purification, while HRV 3C and SUMOstar are detergent-sensitive, and TEV shows intermediate tolerance. These properties directly influence the suitability of each enzyme in practical purification workflows, especially for membrane proteins.
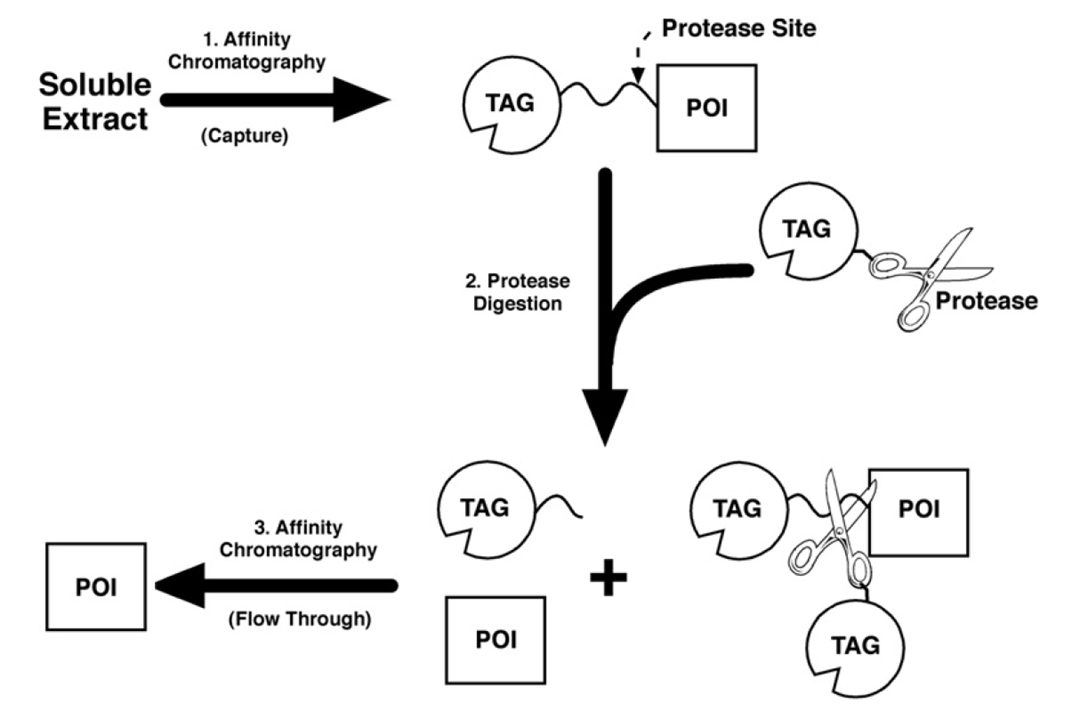
Figure 1. General Strategies for Protein Purification Using Affinity Tag-Cleaving Endoproteases
These enzymes used for affinity tag cleavage mainly belong to the clan CA cysteine protease superfamily. Their hallmark is a highly conserved catalytic triad consisting of Cys-His-Asp. The catalytic mechanism is highly consistent: the Cys acts as the nucleophile attacking the carbonyl carbon of the peptide bond to form a tetrahedral intermediate; His functions as a base to accept a proton, while Asp (or Asn) stabilizes the charge on the His side chain. Together, they facilitate proton transfer and hydrolysis of the acyl-enzyme intermediate, ultimately releasing the cleavage products.
The shared advantage of these enzymes is their relatively low k_cat (typically 1–2 orders of magnitude lower than serine proteases), which trades catalytic speed for extremely high substrate specificity, with virtually no reports of non-specific cleavage. Based on substrate recognition patterns, these enzymes can be clearly divided into two major subclasses: sequence-specific viral proteases and structure-specific deSUMOylases.
These enzymes originate from the polyprotein processing of plant potyviruses and adopt a chymotrypsin-like fold with a catalytic cysteine residue.
These enzymes originate from animal picornaviruses. They are structurally homologous to potyviral NIa proteases and require Gln at P1 along with high activity at low temperatures.
Unlike the linear sequence recognition described above, these enzymes recognize the entire three-dimensional folded structure of the SUMO protein and cleave precisely after the Gly-Gly motif at the C-terminus of SUMO, achieving zero extra residues at the N-terminus of the target protein.
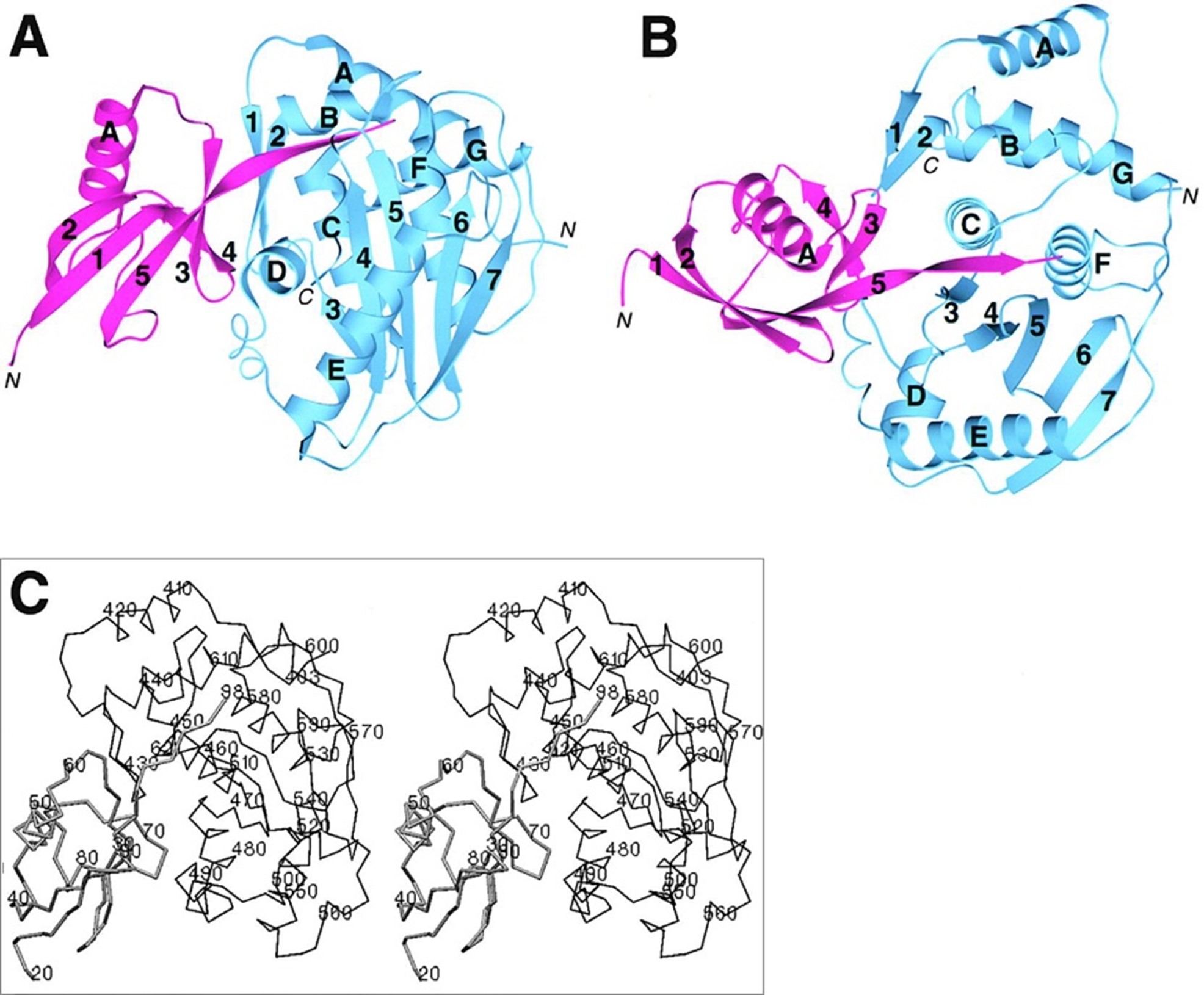
Figure 2. Crystal structure of the Ulp1–SUMO complex (A, B) and detailed view of the active-site pocket (C)
TEV Variant Engineering: The classic S219V mutation has resolved the autolysis issue. Newer variants maintain activity and stability without reducing agents, making them particularly suitable for cleaving proteins containing disulfide bonds.
HRV 3C Optimization: Its low-temperature advantage continues to be validated. Advanced strategies have expanded P1 specificity, offering potential for “scarless” N-terminal cleavage.
SUMO/Ulp1 Advances: SUMOstar (R64T/R71E double mutant) solves the problem of endogenous deSUMOylation in eukaryotic expression systems.
Emerging Enzymes: The high-solubility, high-specificity Con1 protease is expected to complement the limitations of TEV and 3C.
AtaGenix Tag-Cleaving Protease Product Recommendations
| Catalog No. | Product Name |
|---|---|
| ATE00001 | TEV Protease |
| ATE00004 | SUMO Protease |
| ATE00003 | 3C Protease |
Need high-purity, high-activity proteases for recombinant protein tag removal?
AtaGenix offers TEV, 3C, and SUMO proteases with high purity and strong activity to help you obtain near-native target proteins.
Browse All Tag-Cleaving Protease ProductsContact Us
+86-27-6552-3339
info@atagenix.com
Building C, R & D Building, No. 666, Shendun 4th Road, Donghu New Technology Development Zone, Wuhan

